from sklearn.datasets import load_irisDecision Trees
Decision Trees (DTs) are a supervised learning algorithm that learns simple decision rules, to assign predictions to. Example: If age < 30 and income > 50,000 then: "Rich", else: "Not Rich".
Advantages of Decision Trees
Firslty, they can do both Regression and Classification tasks.
Secondly, “White Box”: Simple to understand and to interpret. This is their fundamental advantage. You do not get this level of interpretability with Deep Neural Networks (a “Black Box”)
You can map the exact sequence of logical conditions the algorithm evaluated to reach a conclusion. If a medical triage model recommends prescribing antibiotics, you can trace the decision branches back to see exactly why (e.g., "Body Temperature > 38°C" AND "Rapid Strep Test = Positive").
Trees can also be visualized.
Thirdlay, they can handle Multi-output problems.
Fourthly, inference cost is logarithmic in the number of data points used to train the tree. That means even if your dataset is massive, the time it takes to make a prediction barely increases.
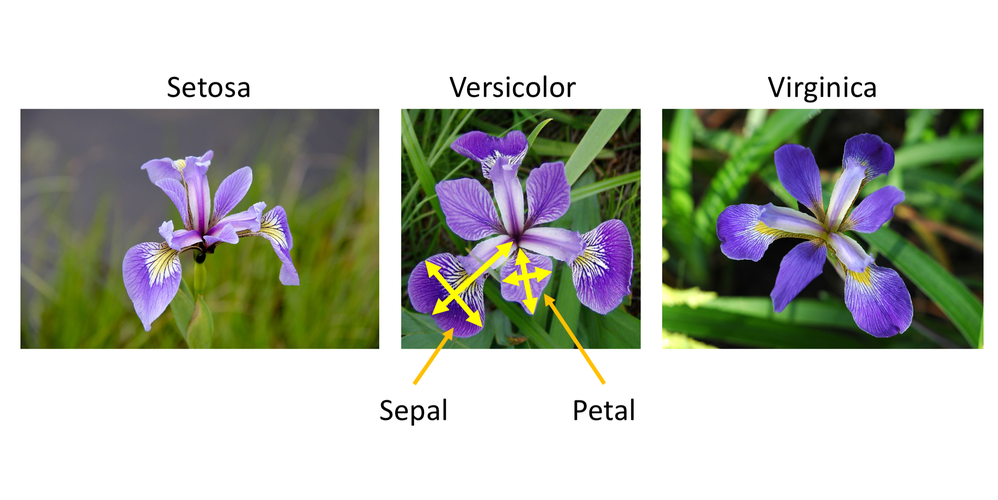
Example: Iris Flowers Classification
For instance, in the example below, decision trees learn from data to approximate a sine curve with a set of if-then-else decision rules. The deeper the tree, the more complex the decision rules and the fitter the model.
iris = load_iris(as_frame=True)
X, y = iris.data, iris.targetimport pandas as pd
pd.concat([iris.data, iris.target], axis=1)| sepal length (cm) | sepal width (cm) | petal length (cm) | petal width (cm) | target | |
|---|---|---|---|---|---|
| 0 | 5.1 | 3.5 | 1.4 | 0.2 | 0 |
| 1 | 4.9 | 3.0 | 1.4 | 0.2 | 0 |
| 2 | 4.7 | 3.2 | 1.3 | 0.2 | 0 |
| 3 | 4.6 | 3.1 | 1.5 | 0.2 | 0 |
| 4 | 5.0 | 3.6 | 1.4 | 0.2 | 0 |
| ... | ... | ... | ... | ... | ... |
| 145 | 6.7 | 3.0 | 5.2 | 2.3 | 2 |
| 146 | 6.3 | 2.5 | 5.0 | 1.9 | 2 |
| 147 | 6.5 | 3.0 | 5.2 | 2.0 | 2 |
| 148 | 6.2 | 3.4 | 5.4 | 2.3 | 2 |
| 149 | 5.9 | 3.0 | 5.1 | 1.8 | 2 |
150 rows × 5 columns
from sklearn.tree import DecisionTreeClassifier
X_iris = iris.data[["petal length (cm)", "petal width (cm)"]].values
y_iris = iris.target
tree_clf = DecisionTreeClassifier(max_depth=2, random_state=42)
tree_clf.fit(X_iris, y_iris)DecisionTreeClassifier(max_depth=2, random_state=42)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
After being fitted, the model can then be used to predict the class of samples:
tree_clf.predict([[5.1, 3.5]])array([2])The probability of each class can be predicted, which is: the fraction of training samples of the class in a leaf:
tree_clf.predict_proba([[5.1, 3.5]]).round(3)array([[0. , 0.022, 0.978]])الكود
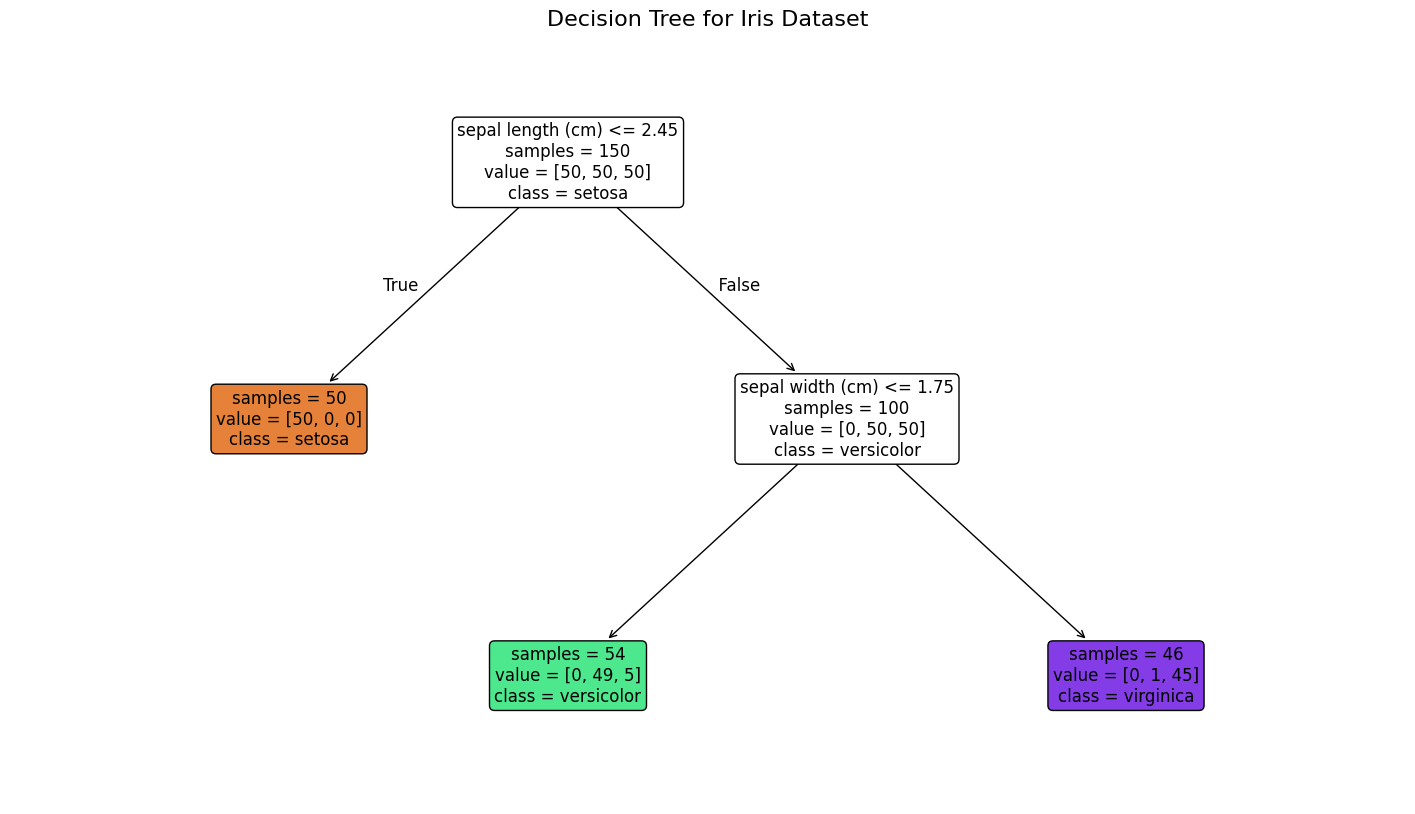
from sklearn.tree import plot_tree
import matplotlib.pyplot as plt
plt.figure(figsize=(18, 10))
plot_tree(
tree_clf,
feature_names=iris.feature_names,
class_names=iris.target_names,
filled=True,
rounded=True,
fontsize=12,
proportion=False,
impurity=False # Hides impurity to reduce clutter
)
plt.title("Decision Tree for Iris Dataset", fontsize=16)
plt.show()الكود
import numpy as np
import matplotlib.pyplot as plt
# extra code – just formatting details
from matplotlib.colors import ListedColormap
custom_cmap = ListedColormap(['#fafab0', '#9898ff', '#a0faa0'])
plt.figure(figsize=(8, 4))
lengths, widths = np.meshgrid(np.linspace(0, 7.2, 100), np.linspace(0, 3, 100))
X_iris_all = np.c_[lengths.ravel(), widths.ravel()]
y_pred = tree_clf.predict(X_iris_all).reshape(lengths.shape)
plt.contourf(lengths, widths, y_pred, alpha=0.3, cmap=custom_cmap)
for idx, (name, style) in enumerate(zip(iris.target_names, ("yo", "bs", "g^"))):
plt.plot(X_iris[:, 0][y_iris == idx], X_iris[:, 1][y_iris == idx],
style, label=f"Iris {name}")
# extra code – this section beautifies and saves Figure 6–2
tree_clf_deeper = DecisionTreeClassifier(max_depth=3, random_state=42)
tree_clf_deeper.fit(X_iris, y_iris)
th0, th1, th2a, th2b = tree_clf_deeper.tree_.threshold[[0, 2, 3, 6]]
plt.xlabel("Petal length (cm)")
plt.ylabel("Petal width (cm)")
plt.plot([th0, th0], [0, 3], "k-", linewidth=2)
plt.plot([th0, 7.2], [th1, th1], "k--", linewidth=2)
plt.plot([th2a, th2a], [0, th1], "k:", linewidth=2)
plt.plot([th2b, th2b], [th1, 3], "k:", linewidth=2)
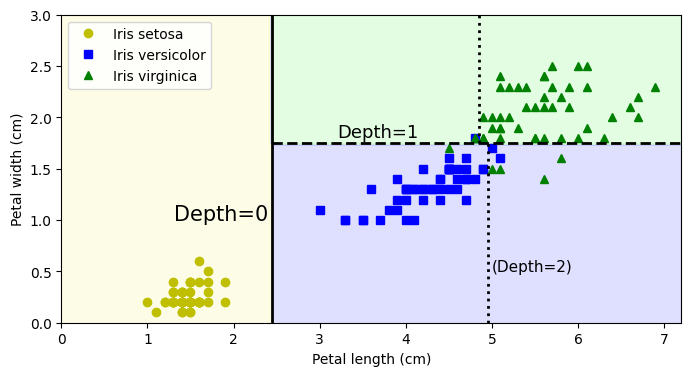
plt.text(th0 - 0.05, 1.0, "Depth=0", horizontalalignment="right", fontsize=15)
plt.text(3.2, th1 + 0.02, "Depth=1", verticalalignment="bottom", fontsize=13)
plt.text(th2a + 0.05, 0.5, "(Depth=2)", fontsize=11)
plt.axis([0, 7.2, 0, 3])
plt.legend()
plt.show()Avoid Overfitting
Decision trees tend to overfit on data with a small samples-to-features ratio. A decision tree will keep growing branches until it has a specific rule for every single data point.
In scikit-learn, restricting a decision tree so it doesn’t memorize your training data is known as regularization or pruning.
If you are just starting out, adjusting max_depth and min_samples_leaf will give you the most immediate control over overfitting.
Let’s illustrate the problem:
from sklearn.datasets import make_moons
X_moons, y_moons = make_moons(n_samples=150, noise=0.2, random_state=42)
tree_clf1 = DecisionTreeClassifier(random_state=42)
tree_clf2 = DecisionTreeClassifier(min_samples_leaf=5, random_state=42)
tree_clf1.fit(X_moons, y_moons)
tree_clf2.fit(X_moons, y_moons)DecisionTreeClassifier(min_samples_leaf=5, random_state=42)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
الكود
def plot_decision_boundary(clf, X, y, axes, cmap):
x1, x2 = np.meshgrid(np.linspace(axes[0], axes[1], 100),
np.linspace(axes[2], axes[3], 100))
X_new = np.c_[x1.ravel(), x2.ravel()]
y_pred = clf.predict(X_new).reshape(x1.shape)
plt.contourf(x1, x2, y_pred, alpha=0.3, cmap=cmap)
plt.contour(x1, x2, y_pred, cmap="Greys", alpha=0.8)
colors = {"Wistia": ["#78785c", "#c47b27"], "Pastel1": ["red", "blue"]}
markers = ("o", "^")
for idx in (0, 1):
plt.plot(X[:, 0][y == idx], X[:, 1][y == idx],
color=colors[cmap][idx], marker=markers[idx], linestyle="none")
plt.axis(axes)
plt.xlabel(r"$x_1$")
plt.ylabel(r"$x_2$", rotation=0)
fig, axes = plt.subplots(ncols=2, figsize=(10, 4), sharey=True)
plt.sca(axes[0])
plot_decision_boundary(tree_clf1, X_moons, y_moons,
axes=[-1.5, 2.4, -1, 1.5], cmap="Wistia")
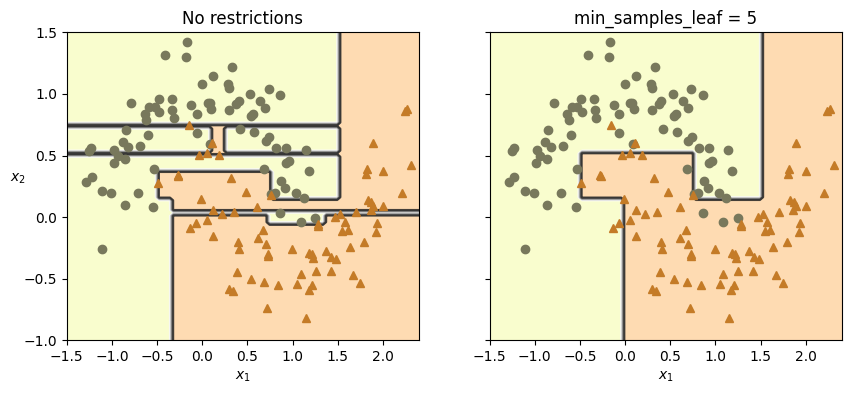
plt.title("No restrictions")
plt.sca(axes[1])
plot_decision_boundary(tree_clf2, X_moons, y_moons,
axes=[-1.5, 2.4, -1, 1.5], cmap="Wistia")
plt.title(f"min_samples_leaf = {tree_clf2.min_samples_leaf}")
plt.ylabel("")
plt.show();X_moons_test, y_moons_test = make_moons(
n_samples=1000,
noise=0.2,
random_state=43
)
print(tree_clf1.score(X_moons_test, y_moons_test))
print(tree_clf2.score(X_moons_test, y_moons_test))0.898
0.921. The Manual Approach: Setting Limits Directly
from sklearn.tree import DecisionTreeClassifier
# Create the tree and set your boundaries right from the start
# We cap the tree at 5 questions deep and force at least 10 data points per final leaf.
my_tree = DecisionTreeClassifier(
max_depth=5,
min_samples_leaf=10
)
# Train the model on your data (X contains your data, y contains the answers)
# my_tree.fit(X, y)2. The Automated Approach: Grid Search
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import GridSearchCV
# 1. Create a blank tree with no rules yet
blank_tree = DecisionTreeClassifier()
# 2. Create your "menu" of options to test (this is your grid)
options_to_test = {
'max_depth': [3, 5, 7, 10, None], # Test shallow, medium, deep, and unrestricted
'min_samples_leaf': [2, 5, 10, 20] # Test different bucket sizes
}
# 3. Set up the automated tester
# cv=5 means "Cross-Validation". It tests every combination 5 separate times
# on different slices of your data to ensure the results aren't just a lucky fluke.
automated_tester = GridSearchCV(
estimator=blank_tree,
param_grid=options_to_test,
cv=5
)
# 4. Start the test (this might take a moment depending on how big your data and menu are)
# automated_tester.fit(X, y)
# 5. Print out the winning combination!
# print("The best limits to use are:", automated_tester.best_params_)Here is the full list of the hyperparameters you can set to stop this from happening, broken down by how they control the tree:
1. Controlling the Architecture (Pre-pruning)
These parameters stop the tree from growing too large in the first place.
max_depth
- What it does: Sets a hard limit on how many levels (or questions) deep the tree can go.
- How it helps: By changing this from its default (
None) to a specific number (like 3, 5, or 10), you force the algorithm to stop drilling down into hyper-specific patterns. It keeps the model focused on the broad, general rules.
min_samples_split
- What it does: Dictates the minimum number of data points a branch must have before it is allowed to split again.
- How it helps: If you set this to 20, any “bucket” of data containing 19 or fewer samples will refuse to split further. This stops the tree from creating complex rules just to satisfy a tiny handful of data points.
min_samples_leaf
- What it does: Dictates the minimum number of data points that must end up in a final leaf node after a split.
- How it helps: A split will only be allowed if both the resulting left and right branches contain at least this many samples. This is highly effective at preventing the tree from isolating random outliers into their own custom leaves.
max_leaf_nodes
- What it does: Caps the total number of endpoints the tree is allowed to have.
- How it helps: If set to 15, the algorithm will find the 15 most important splits (the ones that reduce the most error) and then strictly stop.
2. Introducing Randomness
max_features
- What it does: Limits how many variables (columns) the algorithm is allowed to look at when deciding how to split a node.
- How it helps: If you have 20 variables, you can tell the tree to only consider a random subset of 5 variables at every split. This forces the tree to look for alternative patterns and prevents it from becoming overly dependent on just one dominant variable.
3. Mathematical Pruning (Post-pruning)
ccp_alpha (Cost-Complexity Pruning)
- What it does: It allows the tree to grow fully, but then mathematically calculates the “cost” of each branch. It penalizes branches that are overly complex but don’t add much predictive value.
- How it helps: Increasing this number acts like a pair of hedge clippers. A higher
ccp_alphasnips off weak branches from the bottom up, leaving you with a simpler, more robust tree.
See Example: Post pruning decision trees with cost complexity pruning.
Learn more Tips on Practical Use.
References: https://github.com/ageron/handson-ml3/blob/main/06_decision_trees.ipynb